 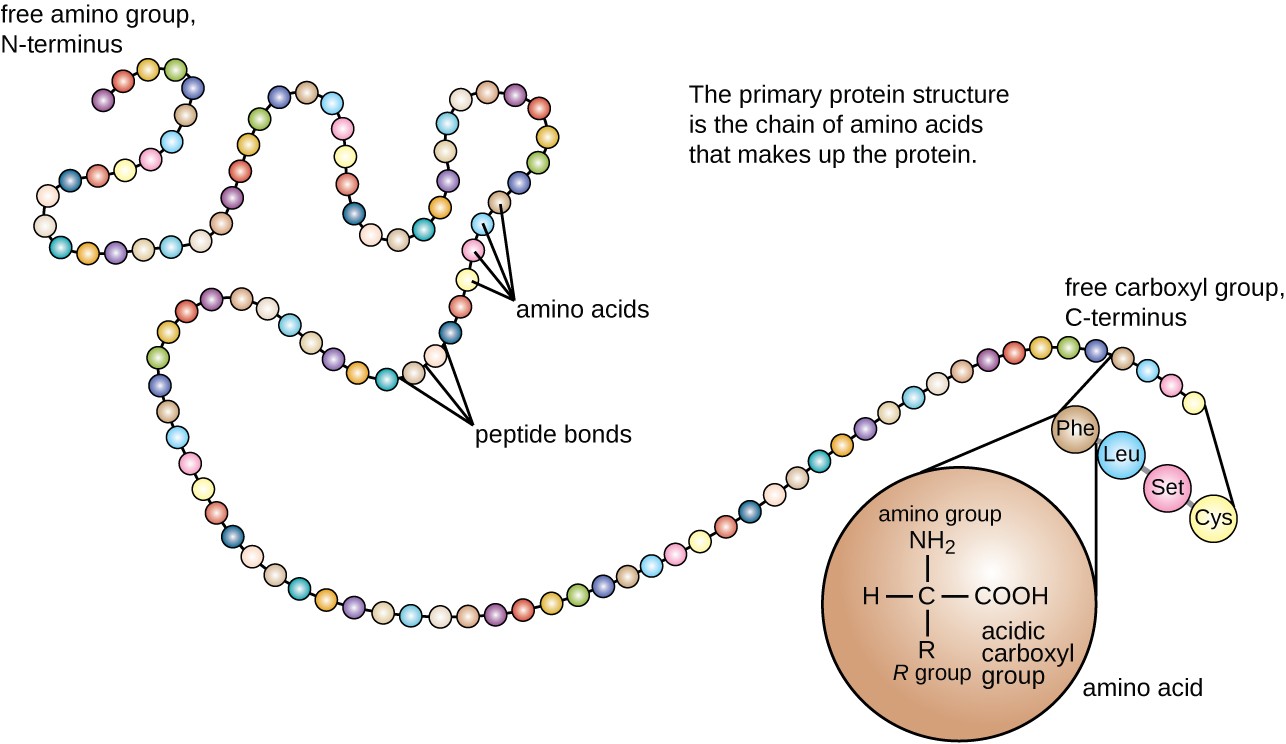 
Extend b-sheet until 4 continuous residues an have an average P(E) 105 and the average P(E) > average P(H) then “b-sheet” Structure Prediction Methods Homology modeling High sequence similarity (> 30% identity) Exploit known whole structure Fold Recognition Medium sequence similarity (generally 100 “alpha-helix nucleus”Įxtend a-helix nucleus 3.Ğxtend helix in both directions until a set of four residues have an average P(H) 100 “b-sheet nucleus” 5. 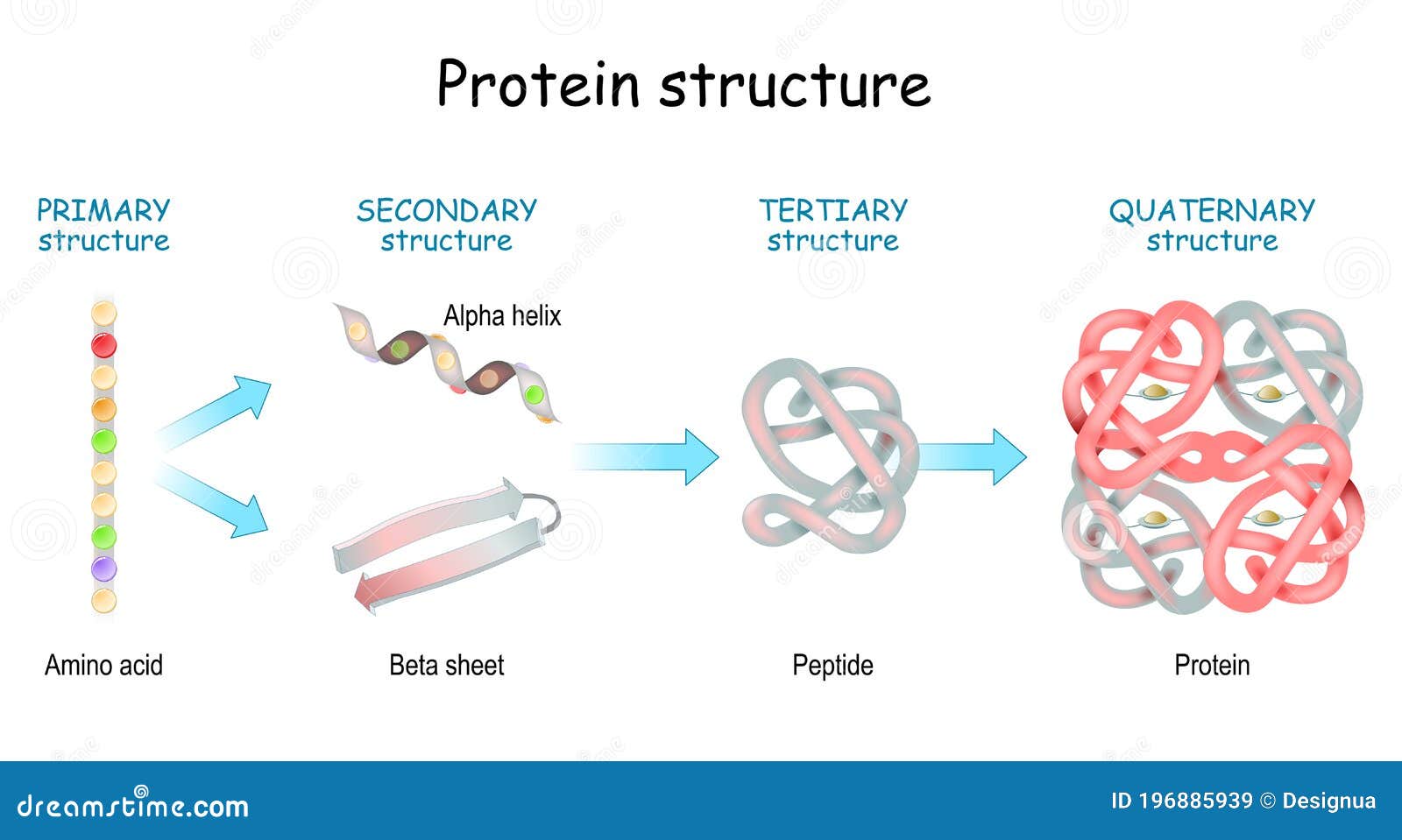
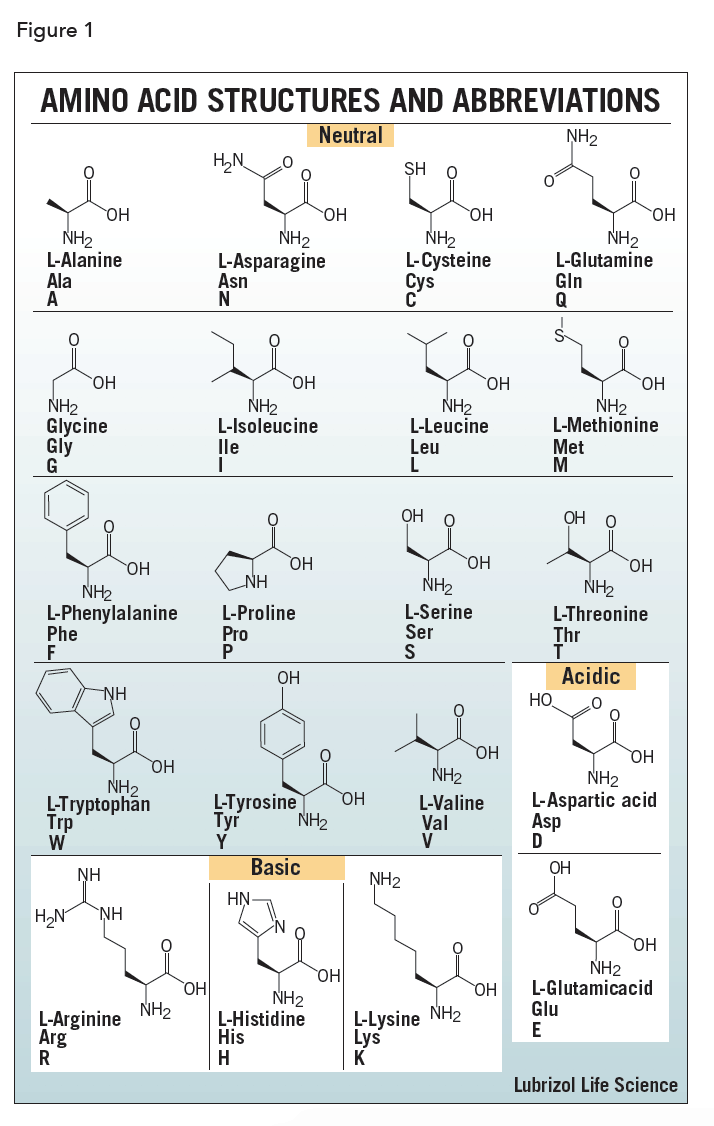
Protein Structure & Function structure medicine Most functions depend on structures sequence function There are only a few thousand possible folds.Įxamples of fold classes (CATH architectures) The core 3D structure of a domain is called a fold.A discrete portion of a protein assumed to fold independently of the rest of the protein and possessing its own function.SECONDARY STRUCTURE (helices, strands) PRIMARY STRUCTURE (amino acid sequence) VHLTPEEKSAVTALWGKVNVDEVGGEALGRLLVVYPWTQRFFESFGDLSTPDAVMGNPKVKAHGKKVLGAFSDGLAHLDNLKGTFATLSELHCDKLHVDPENFRLLGNVLVCVLAHHFGKEFTPPVQAAYQKVVAGVANALAHKYH TERTIARY STRUCTURE (fold) QUATERNARY STRUCTURE (oligomers) Protein structure hierarchical levels -helix -sheet loop/coil (Adapted from Jaap Heringa’s slide) Gene network finding (time series microarray data). 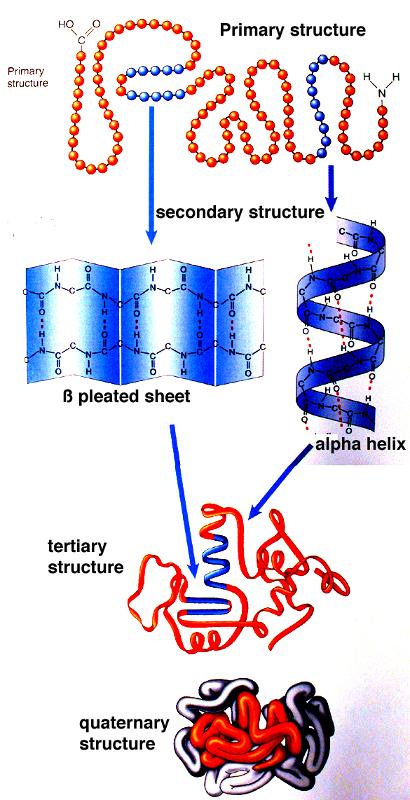
Determine protein-protein interactions.Computational: Profile HMMs, protein classification, motif analysis.Experimental: X-ray Crystallography, Nuclear Magnetic Resonance spectroscopy (NMR), Electron Microscopy/Diffraction.Computational: Sequence matching, energy minimization etc.Determine 3-D protein structures (secondary, tertiary, quaternary).Indirect: Find genes and then translate them to proteins.Determine protein sequences (primary structure).Topics in Bioinformatics Function (Protein) Gene (DNA) Gene expression & regulation Microarray data (Matrix) > DNA sequence AATTCATGAAAATCGTATACTGGTCTGGTACCGGC TGAGAAAATGGCAGAGCTCATCGCTAAAGGTA TCTGGTAAAGACGTCAACACCATCAACGTGTC ACATCGATGAACTGCTGAACGAAGATATCCTG TTGCTCTGCCATGGGCGATGAAGTTCTCGAGG > Protein sequence MKIVYWSGTGNTEKMAELIAKGIIESGKDV DELLNEDILILGCSAMGDEVLEESEFEPFIE KVALFGSYGWGDGKWMRDFEERMNGYG PDEAEQDCIEFGKKIANI transcriptomics Proteomics Genomics Protein Structure Prediction (Lecture for CS397-CXZ Algorithms in Bioinformatics) ApChengXiang Zhai Department of Computer Science University of Illinois, Urbana-Champaign 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
